A robust sequencing assay of a thousand amplicons for the high-throughput population monitoring of Alpine ibex immunogenetics | bioRxiv

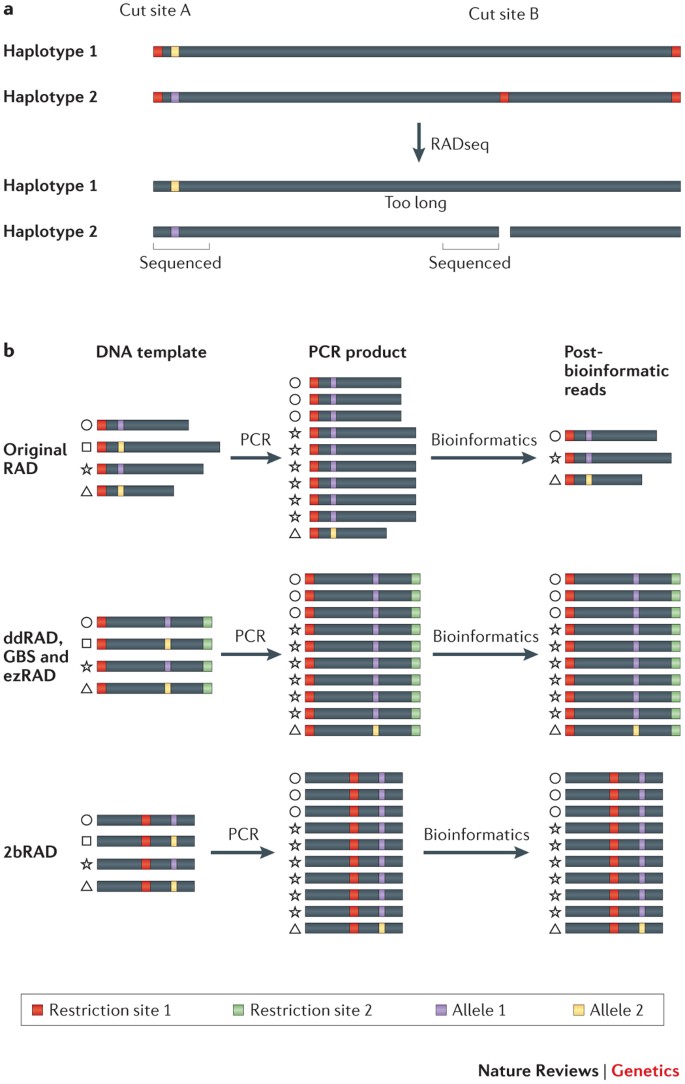
Bioinformatic processing of RAD‐seq data dramatically impacts downstream population genetic inference - Shafer - 2017 - Methods in Ecology and Evolution - Wiley Online Library

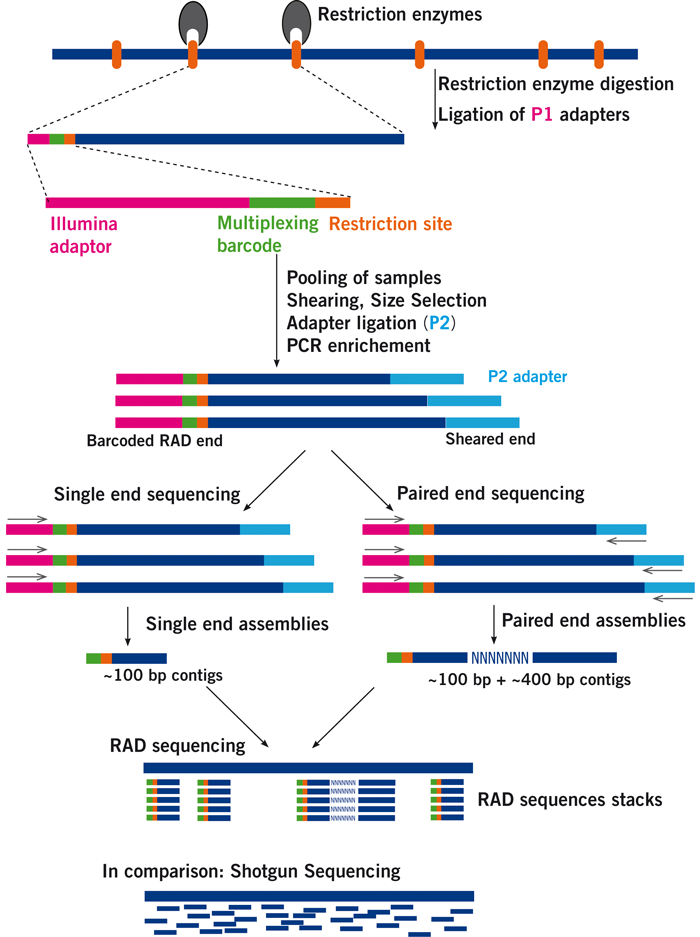
Figure 1 from Paired-end RAD-seq for de novo assembly and marker design without available reference | Semantic Scholar
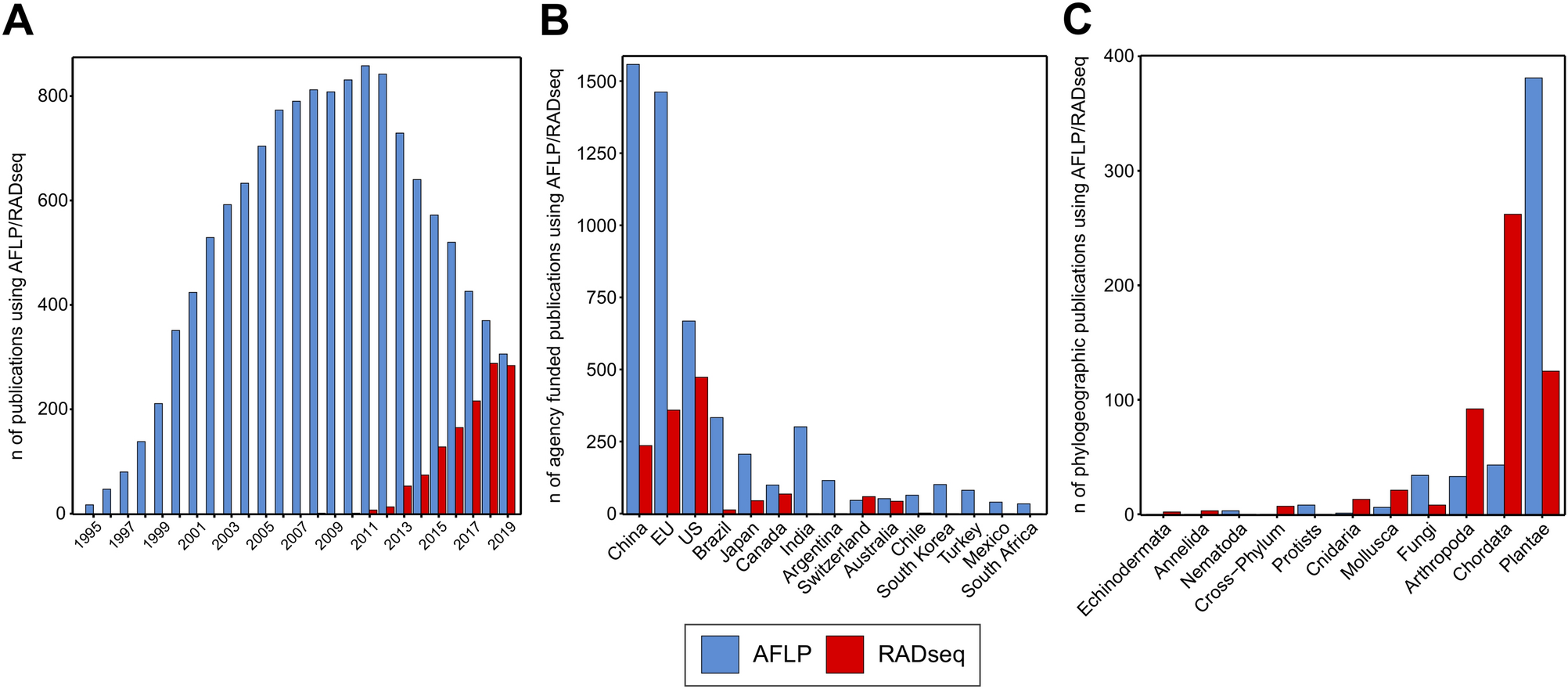
Performance comparison of two reduced-representation based genome-wide marker-discovery strategies in a multi-taxon phylogeographic framework | Scientific Reports

Genome-wide genetic diversity detection and population structure analysis in sweetpotato (Ipomoea batatas) using RAD-seq - ScienceDirect

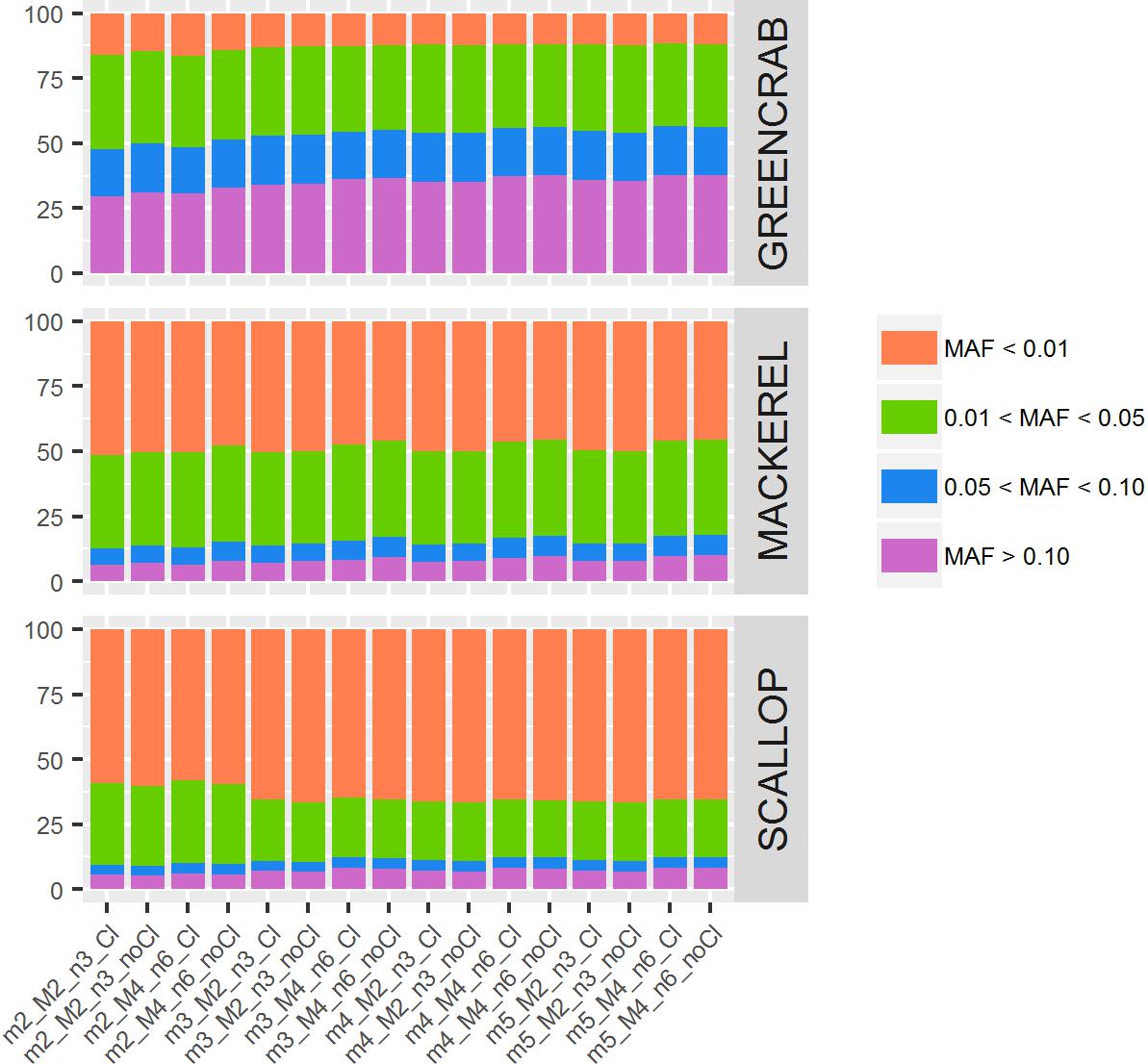
Inherent population structure determines the importance of filtering parameters for reduced representation sequencing analyses | bioRxiv

A combined RAD-Seq and WGS approach reveals the genomic basis of yellow color variation in bumble bee Bombus terrestris | bioRxiv
![dDocent: a RADseq, variant-calling pipeline designed for population genomics of non-model organisms [PeerJ] dDocent: a RADseq, variant-calling pipeline designed for population genomics of non-model organisms [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2014/431/1/fig-1-2x.jpg)
dDocent: a RADseq, variant-calling pipeline designed for population genomics of non-model organisms [PeerJ]
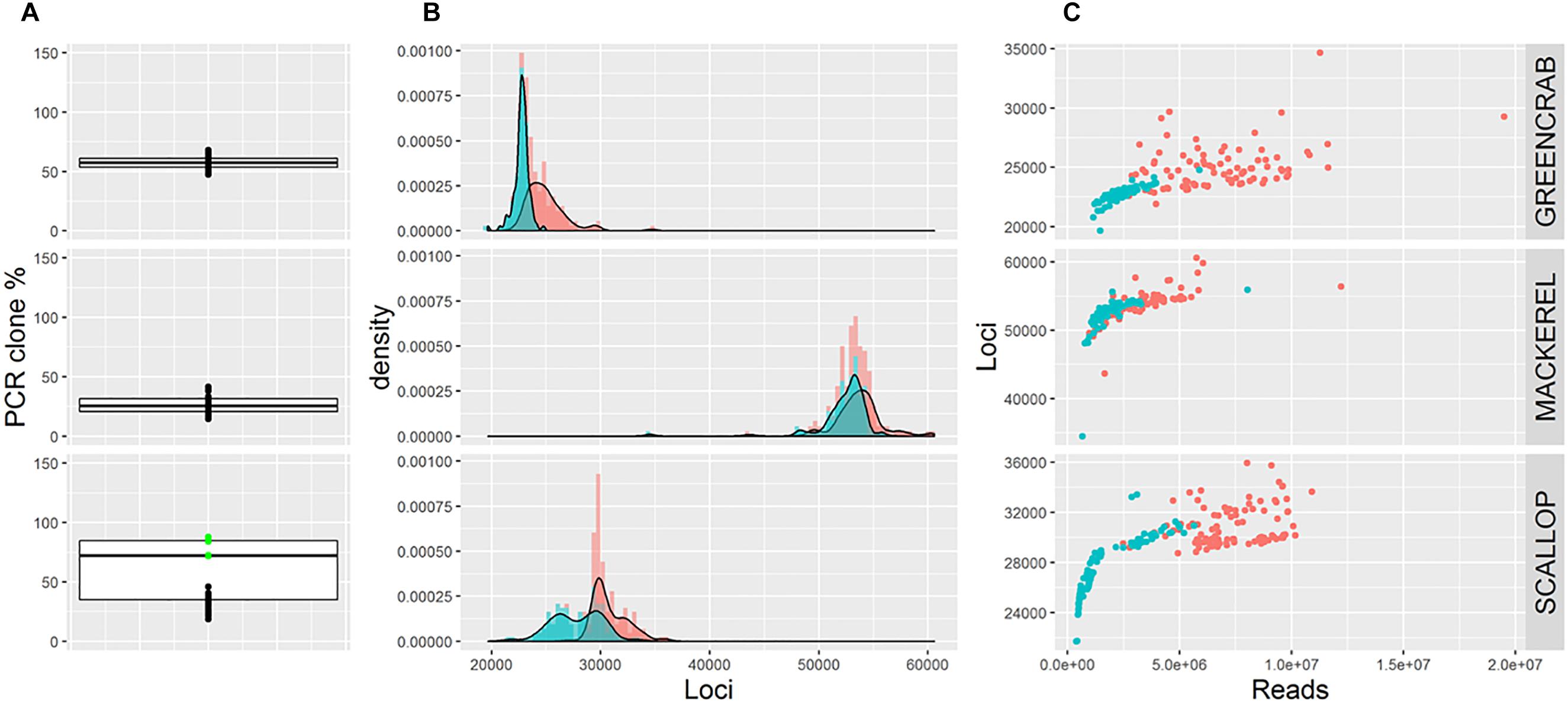
Frontiers | Selecting RAD-Seq Data Analysis Parameters for Population Genetics: The More the Better? | Genetics
Genetic diversity and population structure of the Mediterranean sesame core collection with use of genome-wide SNPs developed by double digest RAD-Seq | PLOS ONE